This is the third in my series of three posts on my package freqparcoord with Yingkang Xie. (My next post after this will show how to use R to explore one of my favorite examples of “what can go wrong” in statistics.)
Here is a very brief review of my previous posts regarding freqparcoord. A parallel coordinates plot draws one vertical axis for each variable, and then draws each data point as a segmented line connecting the values of the variables for that data point. This typically results in heavy screen clutter, but freqparcoord solves the problem by plotting only the data points with the largest estimated multivariate density value.
The package can also be used for outlier hunting (it plots the points with the smallest density values) and cluster hunting (the points corresponding to local maxima of the density are plotted).
In this posting, I will demonstrate yet another use for the package: regression diagnostics. Note that the term regression here means the conditional mean, so classification problems, e.g. logistic regression, are covered too. Specifically, the package includes a function regdiag() that is used to assess fit of a parametric model, using the main function of the package, freqparcoord().
The basic idea is this: The function computes nonparametric estimates of the regression function at all data points, and then computes the difference between the parametric and nonparametric fits at each point; we call those differences the divergences. The latter become the first vertical axis, and then each of the predictor variables has its own vertical axis. The resulting plots then give the user a picture of where in the predictor space the parametric model often overpredicts, and where it often underpredicts. (Note the role of the nonparametric estimates, so that the divergences are NOT the residuals, and note that we are only plotting the “most typical” values of the divergences, due to the use of density by freqparcoord.)
In addition, in the case in which we make use of the output of R’s lm() linear regression function, regdiag() also makes available the adjusted R2 value for the linear model and a corresponding value (squared correlation of actual Y and predicted Y) for the nonparametric fit. If the nonparametric fit is substantially better here, it is an indication that our parametric model may be “leaving money on the table,” i.e. there is room for improvement, say by adding second-order terms.
We use k-NN for the nonparametric fit, by the way, and the estimate at a point uses only genuine neighbors, i.e. the point itself is excluded. Future versions may use a local-linear approximation, in order to deal with aliasing effects. All data is centered and scaled.
Here an example, using the prgeng data set from the package. This is 2000 Census data for programmers and engineers in Silicon Valley. The R code is
data(prgeng) pg <- prgeng pg1 <- pg[pg$wageinc >= 40000 & pg$wkswrkd >= 48,] pg1$ms <- as.integer(pg1$educ == 14) pg1$phd <- as.integer(pg1$educ == 16) pg1$csee <- as.integer(pg1$occ == 140 | pg1$occ == 141) pg1$msphd <- pg1$ms + pg1$phd l1 <- lm(wageinc ~ age+msphd+csee+sex,data=pg1) p <- regdiag(l1,tail=0.40) p p$paramr2 p$nonparamr2
What does this code do? It: loads the data; screens out the odd cases; creates dummy variables for MS/PhD degree and CS/EE job; runs lm(); plugs the output of lm() into regdiag(); “prints” the graph on the screen; and prints out the parametric and nonparametric R2 values.
The latter values are 0.055 and 0.062. These are both low, but the point here is that it does appear that there is room for improvement in the parametric model, possibly by adding second-order terms to the model. Well, then, which second-order terms? The graph gives us some insight here:
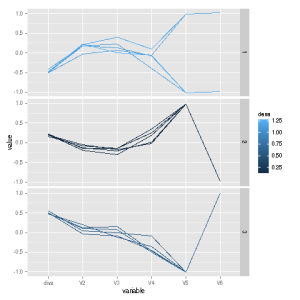
The regdiag() function plots the typical large negative divergences on top, the typical large positive ones on the bottom, and the middling ones in the middle. We see a sharp difference in divergences between the top and bottom plots for the Age variable, but not for the others. This suggests adding an Age2 term to the parametric model, but not adding any interaction terms. After doing this (not shown here), the adjusted R2 value for the parametric model becomes comparable to the nonparametric one.
Let’s do a simulation, so we can see how this all works in a known setting:
n <- 10000 x <- matrix(rnorm(3*n),ncol=3) x <- cbind(x,sample(0:1,n,replace=T)) x <- cbind(x,sample(0:1,n,replace=T)) proddumms <- x[,4] * x[,5] x <- cbind(x,proddumms) mx <- x %*% rep(1,6) y <- mx + rnorm(n,sd=1) l1 <- lm(y ~ x[,1:5]) p <- regdiag(l1,tail=0.40) p
The last two predictors in the original data are dummies, and we add a product term for interaction. We then run lm() without the product term, pretending we don’t know about it, to see if regdiag() will notice. Indeed it does:
The dummy variables are V5 and V6, and the plot indeed shows a lot of activity there! Thus the plot indicates a need for adding an interaction term, as it should.

It works great. Not everybody understands the density thing on the y-axis, but you can pass the idxs to the groupColumn of ggparcoord of the GGally package. That function also allows for plotting on outlyingness and minmax scaling.
Thanks, but I’m not sure I entirely understand your suggestion. Could you explain it using one of the examples in the package?